Now we know more about the deadly diarrhoeal disease that affected dogs in Norway in 2019
New research reveals how dangerous variants of a little-known bacterium differ from harmless ones – and why certain variants in particular can make dogs ill.
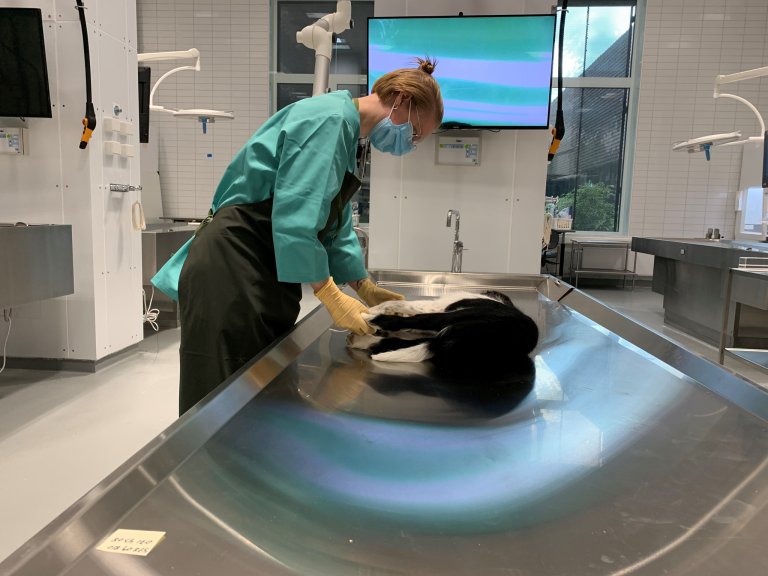

In the autumn of 2019, many dog owners in Norway became frightened. Across the country, dogs suddenly arrived with acute bloody diarrhoea, and several died. Veterinarians were left without clear answers. What was going on?
Acute bloody diarrhoea in dogs can have several causes, and in many cases, no clear explanation is found. This time, however, one bacterium in particular – Providencia alcalifaciens – stood out as the main “suspect” during the investigation. The Norwegian Veterinary Institute, NMBU, and the Norwegian Food Safety Authority launched a comprehensive outbreak investigation, in which this bacterium was detected in a majority of the dogs examined post-mortem and in the faeces of sick dogs. At the time, little was known about its actual role in causing severe intestinal damage in dogs.
Back then, the conclusion was that it was highly likely that Providencia alcalifaciens was involved in the disease, but further research on the collected material was needed to determine whether the bacterium could cause fatal intestinal injury in dogs.
New samples provide new answers
Researchers have made new and important discoveries about this bacterium in a collaborative project involving the Norwegian Veterinary Institute, the NMBU Faculty of Veterinary Medicine, and others. The work has been published in the scientific journal Microbial Genomics.
Veterinarians and animal clinics send samples from dogs with diarrhoea to the Norwegian Veterinary Institute, and in the first years after the major outbreak, the Institute found several cases of Providencia alcalifaciens in late summer and early autumn.
– It had more the character of isolated cases than an outbreak. After that, it seemed to stop, says Bjarne Bergsjø, who is a veterinarian and head of the bacteriology laboratory at the Norwegian Veterinary Institute. All bacterial isolates were preserved for future analyses.
In this study, researchers examined isolates from dogs with diarrhoea collected over several years (2019, 2020 and 2021), in addition to samples from 2005 and 2006*.
An autumn disease
– We found the bacterium only to a small extent outside the autumn season. This observation pointed towards a possible seasonal connection. The finding strengthened our suspicion that the bacterium could have been an important contributing factor in the outbreak, he says.
%20(6).jpg)
At the same time, researchers observed that even when the bacterium was detected, it did not always lead to illness. This made them ask whether there was something specific about the bacterial strain that caused the outbreak.
Investigating the “family tree”
To understand what made the bacterium dangerous, the researchers analysed samples of P. alcalifaciens collected in Norway (273 in total), the vast majority from Norwegian dogs. These were compared with 48 P. alcalifaciens samples from other studies, including international sources. The team used whole genome sequencing – an analysis that “reads” the entire DNA of the bacteria to determine how they are related.
The analysis revealed a clear pattern. The bacteria fell into two genetic groups: A and B. And the particularly interesting findings were in group A. The investigations showed that bacteria from group A were overrepresented in samples from sick dogs, including many from the 2019 outbreak.
Particularly aggressive types
A closer look at the subgroups within group A showed that some of them frequently carried additional genes associated with the ability to cause disease.
– Several subgroups stood out by having plasmids that resembled one another, says PhD fellow Eiril Moen Soltvedt at the NMBU Faculty of Veterinary Medicine. This multi-year work forms part of her doctoral research.
She explains that plasmids are pieces of DNA that bacteria can exchange – a kind of “downloadable toolbox”. This “package” of genes makes these variants particularly aggressive.
In other bacterial species, similar genes provide the bacterium with a type of syringe used to inject proteins into intestinal cells, causing them to stop functioning normally. The injected proteins can enable the bacteria to break down the intestinal mucus layer and suppress the immune response.
In this study, the researchers found genes resembling those found in dangerous E. coli strains and a toxin gene found in the plague bacterium Yersinia.
– We found that almost all bacteria in subgroups A‑1 and A‑4 carried such ‘toolboxes’ with potential virulence genes. This suggests that these dangerous variants have not evolved slowly, but rather ‘downloaded’ the traits that make them harmful, Soltvedt explains.

What does this mean for dog owners?
Simen Foyn Nørstebø, researcher at the Norwegian Veterinary Institute and co-author of the study, believes the findings have direct practical implications.
–Now that we know that specific variants of P. alcalifaciens appear to emerge each autumn in Norway and are particularly dangerous for dogs, veterinarians can more easily identify this bacterium as the cause. This means that in time we may develop more targeted treatment – and possibly even prevent disease, he says.
Researchers also found the bacterium in some healthy dogs. This suggests that P. alcalifaciens does not always cause disease on its own, but exploits situations in which the dog is weakened – for example by stress, another illness, or intestinal injury.
Previous advice still applies
Some soil samples collected near the Veterinary Hospital at NMBU in 2021 were almost identical to bacteria found in sick dogs.
– Whether this is because the bacterium can survive in soil on its own, or because the soil was contaminated by faeces from sick dogs, is currently unclear, says Nørstebø. Regardless of the cause, the finding suggests that the environment may play a role in the transmission chain.
He adds that advice given during previous outbreaks still applies:
Keep dogs away from rotten food scraps, rubbish, faeces and stagnant water, and contact a veterinarian if your dog develops bloody or severe diarrhoea and shows reduced general condition. Most dogs that receive appropriate veterinary care for such symptoms survive.
He believes that monitoring this bacterium can help us be better prepared when next autumn approaches.
The way forward
The next step is to investigate whether the disease-associated genes are actually active when the bacterium is in the dog’s intestine, and what role they play in disease development. There is also a need to find out why some dogs are more vulnerable than others, and to determine where the bacterium persists in nature between outbreaks.
– The answers to these questions will determine whether we can one day prevent outbreaks rather than simply respond to them, says Nørstebø.
A Norwegian piece in a global puzzle – the same type of bacterium makes both animals and humans ill
The researchers also compared the Norwegian bacteria with those found elsewhere in the world. The comparison shows that the most dangerous variants belong to a clear international lineage, but with a Norwegian “branch” that played a central role in the outbreak here.
– It is interesting to see that bacteria from dogs and humans fell into the same genetic subgroups, says Nørstebø.
– For example, we found a Norwegian dog bacterium that was very similar to a bacterium from a human with diarrhoea. This may mean that the bacterium can jump between species, like several other bacteria that cause disease in both animals and humans.
–In the 2019 outbreak, we did not detect any cases of humans being infected by sick dogs. This suggests that several conditions must be present before such bacteria can infect humans. Nevertheless, it is sensible to maintain good hygiene when handling a sick dog with diarrhoea, Nørstebø concludes.
The study is published in Microbial Genomics:
Canine Providencia alcalifaciens: virulence factors and phylogenetic analysis of an emerging enteropathogen | Microbiology Society
*About a quarter of the samples were collected at the NMBU Faculty of Veterinary Medicine, and the remaining samples at the Norwegian Veterinary Institute.


